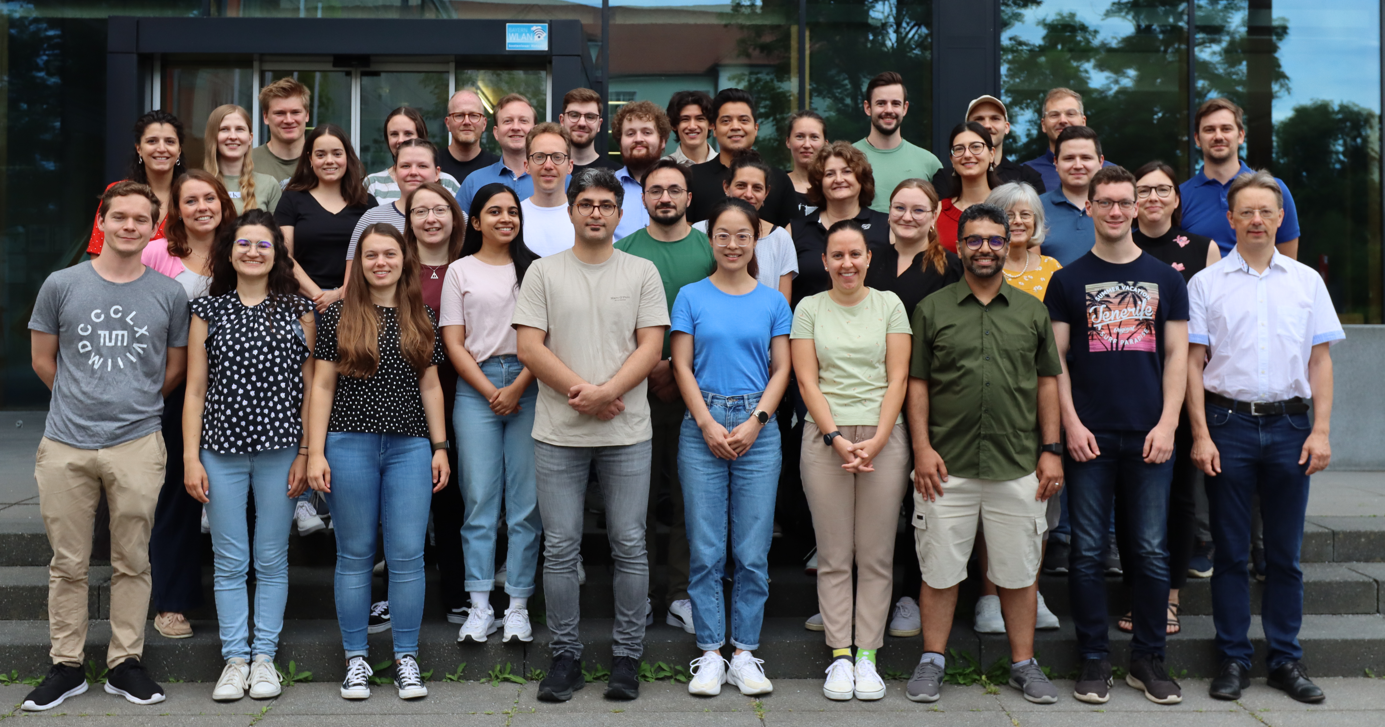
Head

Prof. Dr. Volker Sieber
Chair of Chemistry of Biogenic Resources
- Rector TUMCS
- Head of Chair
- Phone:
- +49 9421 187-300
- Email:
- sieber@tum.de
Office

Anna Kronewald
Chair of Chemistry of Biogenic Resources
- Team Assistant
- Phone:
- +49 9421 187-301
- Email:
- anna.kronewald@tum.de

Eva Groll
Chair of Chemistry of Biogenic Resources
- Team Assistant
- Phone:
- +49 9421 187-307
- Email:
- eva.groll@tum.de
Research Assistants and Doctoral Students

Dr.-Ing. Ammar Al-Shameri
Chair of Chemistry of Biogenic Resources
- Postdoctoral Researcher & Groupleader
- Phone:
- +49 9421 187-322
- Email:
- a.al-shameri@tum.de

Dr. Sara Arana Pena
Chair of Chemistry of Biogenic Resources
- Postdoctoral Researcher, Groupleader
- Phone:
- +49 9421 187-343
- Email:
- sara.arana@tum.de

Dr. Alexander Fuchs
Chair of Chemistry of Biogenic Resources
- Postdoctoral Researcher
- Phone:
- +49 9421 187-320
- Email:
- alex.fuchs@tum.de
Antonio Giustino, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-305
- Email:
- antonio.giustino@tum.de

Jan Grundheber, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-330
- Email:
- jan.grundheber@tum.de

Niklas Hofer, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-323
- Email:
- niklas.hofer@tum.de

Dr. Thomas Kinateder
Chair of Chemistry of Biogenic Resources
- Postdoctoral Researcher
- Phone:
- +49 9421 187-331
- Email:
- Thomas.kinateder@tum.de

Tobias Köllen, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 89 289-54209
- Email:
- tobias.koellen@tum.de
TUM Catalytic Research Center
Ernst-Otto-Fischer-Straße 1
85748 Garching
Germany
Room: 2018

Viktoria Lehmann, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-325
- Email:
- viktoria.lehmann@tum.de

Dr. Yajing Liu
Chair of Chemistry of Biogenic Resources
- Postdoctoral Researcher, Research Assistant
- Phone:
- +49 9421 187-333
- Email:
- yajing.liu@tum.de
Zahabbyia Malubhoy, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-345
- Email:
- z.malubhoy@tum.de
TUM Catalytic Research Center
Ernst-Otto-Fischer-Straße 1
85748 Garching
Germany

Matea Marosevic, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-313
- Email:
- matea.marosevic@tum.de

Franka Matena, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-324

Marcel Mayer, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-316
- Email:
- ma.mayer@tum.de

Tobias Ostertag, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-315
- Email:
- tobias.ostertag@tum.de

Dr. Enrique Raga Carbajal
Chair of Chemistry of Biogenic Resources
- Postdoctoral Researcher, Groupleader
- Phone:
- +49 9421 187-306
- Email:
- enrique.raga@tum.de

Maximiliane Rau, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-287
- Email:
- maximiliane.rau@tum.de

Dennis Romeis, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-178
- Email:
- dennis.romeis@tum.de

Dr. Broder Rühmann
Chair of Chemistry of Biogenic Resources
- Postdoc & Groupleader
- Phone:
- +49 9421 187-332
- Email:
- broder.ruehmann@tum.de
Jonathan Scheerer, M.Sc.
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 9421 187-349
- Email:
- jonathan.scheerer@tum.de

Dr. Doris Schieder
Chair of Chemistry of Biogenic Resources
- Postdoc & Groupleader
- Phone:
- +49 9421 187-303
- Email:
- doris.schieder@tum.de

Dr. Amelie Skopp
Chair of Chemistry of Biogenic Resources
- Research Fellow
- Groupleader
- Phone:
- +49 9421 187-345
- Email:
- amelie.skopp@tum.de
TUM Campus Straubing
Schulgasse 16
94315 Straubing

Lianqing Tong
Chair of Chemistry of Biogenic Resources
- Doctoral Candidate
- Phone:
- +49 89 289-54209
- Email:
- l.tong@tum.de
TUM Catalytic Research Center
Ernst-Otto-Fischer-Straße 1
85748 Garching
Germany
Room: 2018
Technical Personnel

Dipl.-Ing. (FH) Manuel Döring
Chair of Chemistry of Biogenic Resources
- Technical Assistant
- Phone:
- +49 9421 187-319
- Email:
- manuel.doering@tum.de

Magdalena Haslbeck
Chair of Chemistry of Biogenic Resources
- Technical Assistant
- Phone:
- +49 9421 187-327
- Email:
- m.haslbeck@tum.de

Dipl.-Ing. (FH) Anja Schmidt
Chair of Chemistry of Biogenic Resources
- Technical Assistant
- Phone:
- +49 9421 187-336
- Email:
- a.v.schmidt@tum.de

Kai Stirnweiß
Chair of Chemistry of Biogenic Resources
- Technical Assistant
- Phone:
- +49 9421 187-337
- Email:
- kai.stirnweiss@tum.de

Lena Würstl
Chair of Chemistry of Biogenic Resources
- Technical Asistant
- Phone:
- +49 9421 187-326
- Email:
- lena.wuerstl@tum.de
Alumni
| Prof. Dr.-Ing. Jochen Schmid |
| Daniel Bauer |
| Dominik Schwarz |
| Dr. Herbert Goldinger |
| Dr. Claudia Nowak |
| Dr. Steven Koenig |
| Dr. Tobias Gmelch |
| Dr. Nadine Sperl |
| Dr. Michael Loscar |
| Dr. Jose Guillermo Ortiz Tena |
| Dr. Ulrike Obst |
| Dr. Steffen Roth |
| Janine Simon |
| Lisa Steiner |
| Janine Huber |
| Karola Wiesmüller |
| Dr. Fabian Steffler |
| Dr. Marius Rütering |
| Dr. Irina Funk |
| Dr. Hendrik Hohagen |
| Alfiya Wohn |
| Andre Pick |
| Qianqin Zhu |
| Dr. Nina Rimmel |
| Dr. Nadine Borst |
| Dr. Jörg Carsten |
| Ioannis Zachos |
| Dr. Paul Stockmann |
| Dr.-Ing. Sumanth Ranganathan |
| Scott Bottoms |
| Dr. Joseph Sperl |
| Dr. Edilberto Medina Cabrera |
| Dr. Moritz Gansbiller |
| Dr. Robert Genth |
| Dr. Samed Güner |
| Dr. Korbinian Sinzinger |
| Dr. Barbara Beer |
| Dr. Benjamin Begander |
| Dr. Samuel Sutiono |
| Dr. Christoph Schilling |
| Dr. Torben Hüsing |
| Dr. Johanna Radomski |
| Mathilde Steinmaßl |
| Dr. Okke Melse |
| Matthias Steiger |
| Tatjana Laudage |
| Alexander Benson |
| Dr.-Ing. Tristan Rath |
| Dr. Luca Schmermund |
| Dr. Vivian Willers |
| Dr. Mariko Teshima |
| Masoud Babakhani |
| Ernesto Rodriguez |
| Eleonora Fornoni |
| Dr. Enrico Hupfeld |
| Amir Hossein Abbas Nia |
| Zinnia Dsouza |
| Jan-Simon Friedrichs |
| Dominik Siebert |
| Mayla Schulz |
| Kathrin Hörnschemeyer |
- Prof. Dr.-Ing. Jochen Schmid
- Daniel Bauer
- Dominik Schwarz
- Dr. Herbert Goldinger
- Dr. Claudia Nowak
- Dr. Steven Koenig
- Dr. Tobias Gmelch
- Dr. Nadine Sperl
- Dr. Michael Loscar
- Dr. Jose Guillermo Ortiz Tena
- Dr. Ulrike Obst
- Dr. Steffen Roth
- Janine Simon
- Lisa Steiner
- Janine Huber
- Karola Wiesmüller
- Dr. Fabian Steffler
- Dr. Marius Rütering
- Dr. Irina Funk
- Dr. Hendrik Hohagen
- Alfiya Wohn
- Andre Pick
- Qianqin Zhu
- Dr. Nina Rimmel
- Dr. Nadine Borst
- Dr. Jörg Carsten
- Ioannis Zachos
- Dr. Paul Stockmann
- Dr.-Ing. Sumanth Ranganathan
- Scott Bottoms
- Dr. Joseph Sperl
- Dr. Edilberto Medina Cabrera
- Dr. Moritz Gansbiller
- Dr. Robert Genth
- Dr. Samed Güner
- Dr. Korbinian Sinzinger
- Dr. Barbara Beer
- Dr. Benjamin Begander
- Dr. Samuel Sutiono
- Dr. Christoph Schilling
- Dr. Torben Hüsing
- Dr. Johanna Radomski
- Mathilde Steinmaßl
- Dr. Okke Melse
- Matthias Steiger
- Tatjana Laudage
- Alexander Benson
- Dr.-Ing. Tristan Rath
- Dr. Luca Schmermund
- Dr. Vivian Willers
- Dr. Mariko Teshima
- Masoud Babakhani
- Ernesto Rodriguez
- Eleonora Fornoni
- Dr. Enrico Hupfeld
- Amir Hossein Abbas Nia
- Zinnia Dsouza
- Jan-Simon Friedrichs
- Dominik Siebert
- Mayla Schulz
- Kathrin Hörnschemeyer
